

This is an implementation of the funnel plot processes, and
overdispersion methods described in:
Statistical
methods for healthcare regulation: rating, screening and surveillance.
Spiegelhalter et al (2012)
Funnel plots for comparing
institutional performance. Spiegelhalter (2005)
Handling
over-dispersion of performance indicators. Spiegelhalter
(2005)
It draws funnel plots using ggplot2 and allows users to
specify whether they want to adjust the funnel plot limits for
‘overdispersion.’ This adjustment makes the assumption that we are
dealing with clusters of values (means) at institutions that are
themselves arranged around a global mean. We then have ‘within’
institution variation and ‘between institution’ variation. The process
assessed the expected variance in our data, and where it is greater than
that expected by the Poisson distribution, uses the difference as a
scaling factor. It is then used in an additive fashion, after an
adjustment for outliers by either Winsorised or truncated (with a
default 10% at each end of the distribution.)
Methods are based on those presented in Spiegelhalter’s papers and
the Care Quality Commission’s Intelligent Monitoring methodology
documents, with methods for proportions, ratios of counts and indirectly
standardised ratios. There is a also a variant method for standardised
ratios, used in the NHS’ Summary Hospital Mortality Indicator’
Summary
Hospital-level Mortality Indicator, NHS Digital, SHMI specification
This variant uses a log-transformation and truncation of the distribution for calculating overdispersion, whereas Spiegelhalter’s methods use a square-root and Winsorisation.
Contributions are welcome. Please note that the ‘FunnelPlotR’ project is released with a Contributor Code of Conduct. By contributing to this project, you agree to abide by its terms.
More information available at https://nhs-r-community.github.io/FunnelPlotR/
You can install from CRAN:
install.packages("FunnelPlotR")You can install the development version directly from GitHub using
the remotes (or devtools) package. Please be
aware that, although I endeavour have help files up-to-date, this
version may different from the one on CRAN. Please consult the help
documentation if you get error messages.
remotes::install_github("https://github.com/nhs-r-community/FunnelPlotR")We will load the medpar dataset from Hilbe’s
COUNT package. This is based on 1991 Medicare files for the
state of Arizona (Hilbe, Joseph M (2014), Modeling Count Data,
Cambridge University Press). We will first load the data and build
a simple predictive model using a Poisson GLM.
library(FunnelPlotR)
library(COUNT)
library(ggplot2)
# lets use the 'medpar' dataset from the 'COUNT' package. Little reformatting needed
data(medpar)
medpar$provnum<-factor(medpar$provnum)
medpar$los<-as.numeric(medpar$los)
mod<- glm(los ~ hmo + died + age80 + factor(type), family="poisson", data=medpar)
summary(mod)
#>
#> Call:
#> glm(formula = los ~ hmo + died + age80 + factor(type), family = "poisson",
#> data = medpar)
#>
#> Coefficients:
#> Estimate Std. Error z value Pr(>|z|)
#> (Intercept) 2.26875 0.01246 182.011 < 2e-16 ***
#> hmo -0.07637 0.02393 -3.192 0.00142 **
#> died -0.24574 0.01826 -13.458 < 2e-16 ***
#> age80 -0.02141 0.02050 -1.045 0.29617
#> factor(type)2 0.24921 0.02099 11.871 < 2e-16 ***
#> factor(type)3 0.74869 0.02627 28.496 < 2e-16 ***
#> ---
#> Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
#>
#> (Dispersion parameter for poisson family taken to be 1)
#>
#> Null deviance: 8901.1 on 1494 degrees of freedom
#> Residual deviance: 7977.7 on 1489 degrees of freedom
#> AIC: 13705
#>
#> Number of Fisher Scoring iterations: 5Now we have a regression that we can use to get a predicted
los that we will compare to observed los:
medpar$prds <- predict(mod, type="response")
We can build a funnel plot object with standard Poisson
limits, and outliers labelled.
a <- funnel_plot(medpar, numerator = los, denominator = prds, group = provnum,
title = 'Length of Stay Funnel plot for `medpar` data', data_type = "SR", limit= 99,
draw_unadjusted = TRUE, draw_adjusted = FALSE, label = "outlier")
print(a)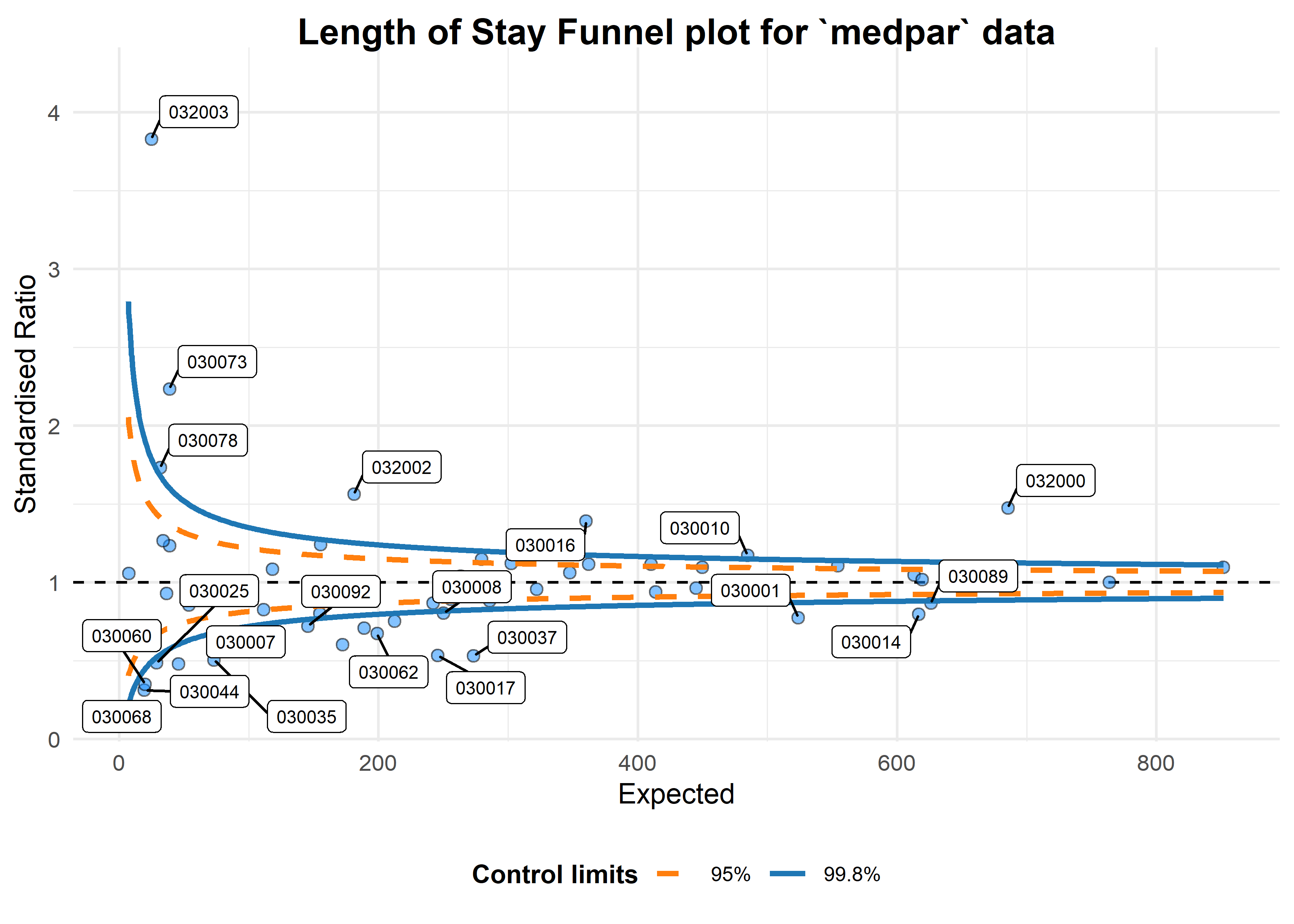
#> A funnel plot object with 54 points of which 25 are outliers.
#> Plot is not adjusted for overdispersion.That looks like too many outliers! There is more variation in our
data than we would expect, and this is referred to as:
overdispersion. So lets check for it:
The
following ratio should be 1 if our data are conforming to Poisson
distribution assumption (conditional mean = variance). If it is greater
than 1, we have overdispersion:
sum(mod$weights * mod$residuals^2)/mod$df.residual
#> [1] 6.240519This suggest the variance is 6.24 times the condition mean, and definitely overdispersed. This is a huge topic, but applying overdispersed limits using either SHMI or Spiegelhalter methods adjust for this by inflating the limits:
b <- funnel_plot(medpar, numerator = los, denominator = prds, group = provnum, data_type = "SR",
title = 'Length of Stay Funnel plot for `medpar` data', draw_unadjusted = FALSE,
draw_adjusted = TRUE, sr_method = "SHMI", label = "outlier", limit = 99)
print(b)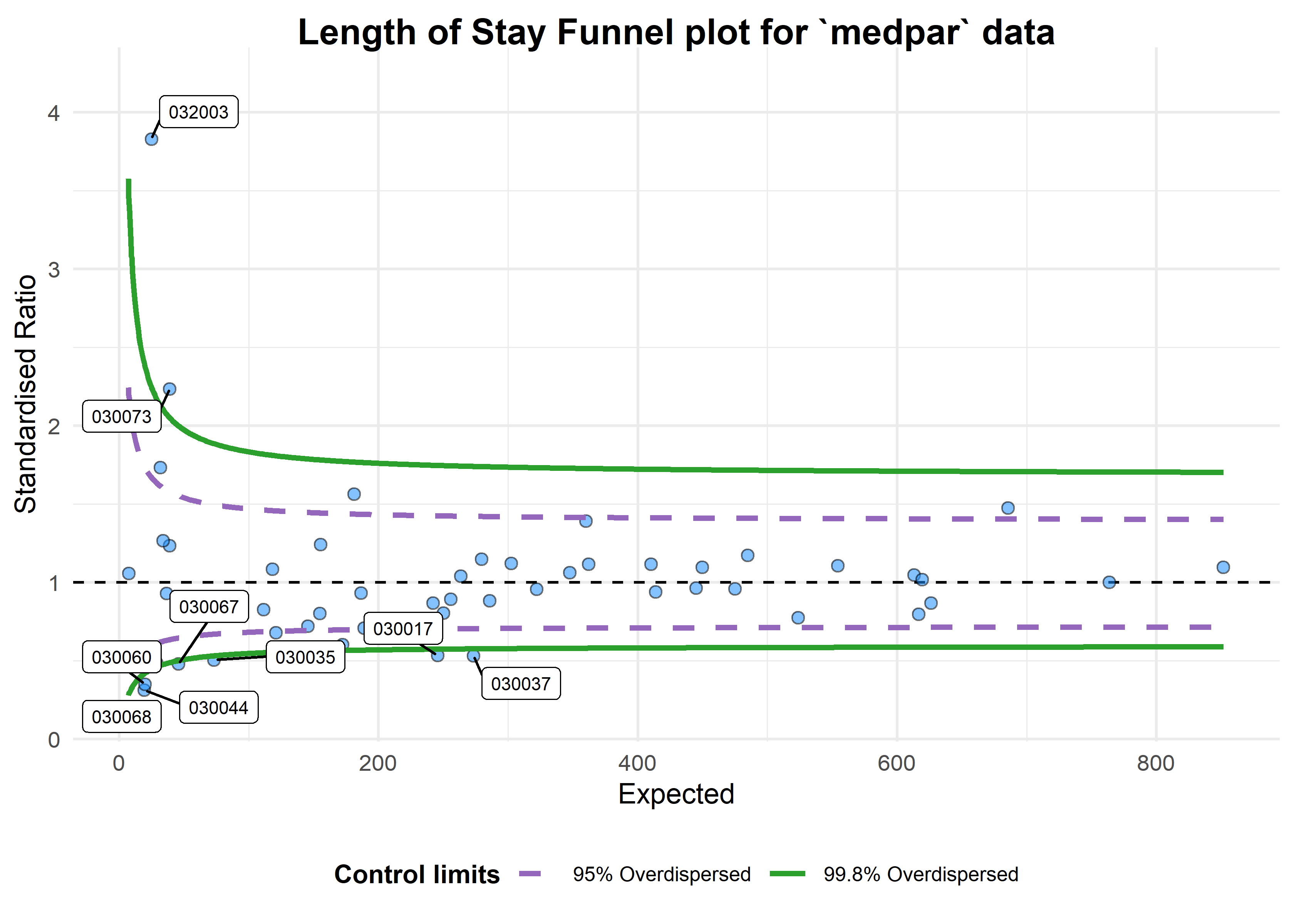
#> A funnel plot object with 54 points of which 9 are outliers.
#> Plot is adjusted for overdispersion.
These methods can be used for any similar indicators,
e.g. standardised mortality ratios, readmissions etc.
Please read the package documentation for more info, at: https://nhs-r-community.github.io/FunnelPlotR/
Funnel Plot HEX sticker/logo by Paul Chipperfield